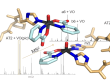
By Tao Jianmei. It is very important to study the protein-binding behavior of platinum drugs in blood. Mass spectrometry-based strategies have been used to identify and quantitate the proteins binding with platinum anticancer drugs in blood, as reviewed by Du et al in Pharmaceuticals. However, early attempts to identify the binding proteins of platinum drugs in serum were restricted by the software for mass spectrometric data analysis. Since the blood proteome is complicated, false-positive results will occur, especially those peptides with the same mass but different elemental compositions as that of platinated peptide. Thus, the proteomic search engine cannot effectively exclude false positive results. Consequently, manually checking the isotope pattern of modified peptides is the only way to exclude false-positive results.
Recently, our recent studies combined the high-abundance protein separation and ion mobility mass spectrometry-based 4D proteomic method to systematically and comprehensively identify the binding proteins of cisplatin in blood. The characteristic isotope patterns of platinated peptides and a similarity algorithm apm2s were utilized to eliminate false-positive identification automatically. Finally, 39 proteins were found to be platinated. The binding ratio of some peptides with cisplatin was measured based on the area ratio of the free peptide using the parallel reaction monitoring method. These results were published in Journal of Proteome Research.

Tao Jianmei

Jia Shuailong