Excercise 1
Charge in Mass Spectrometry
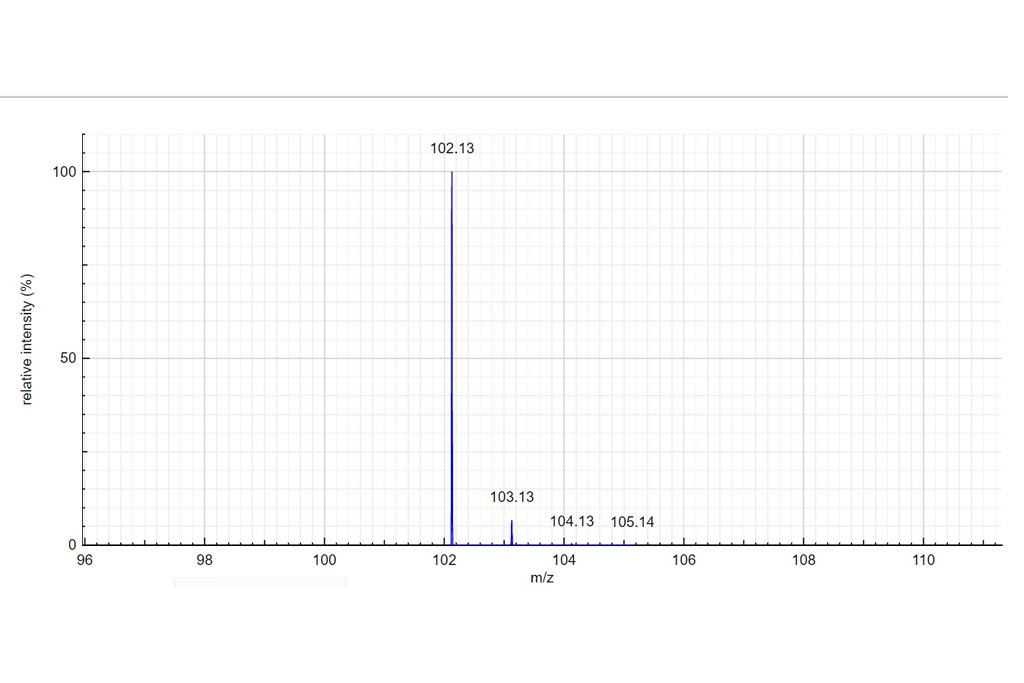
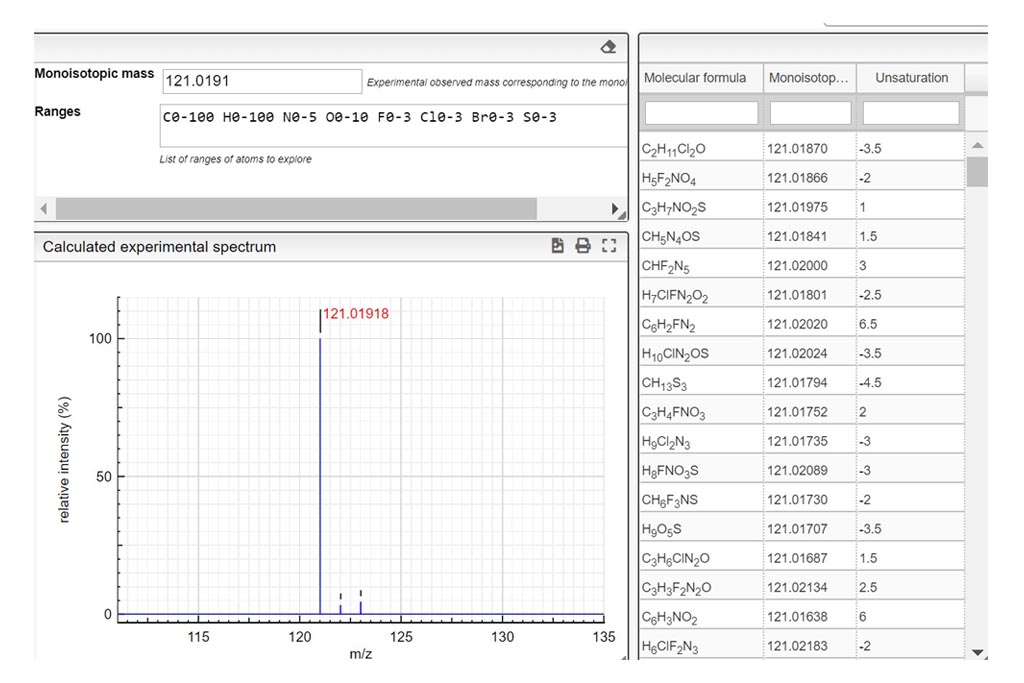
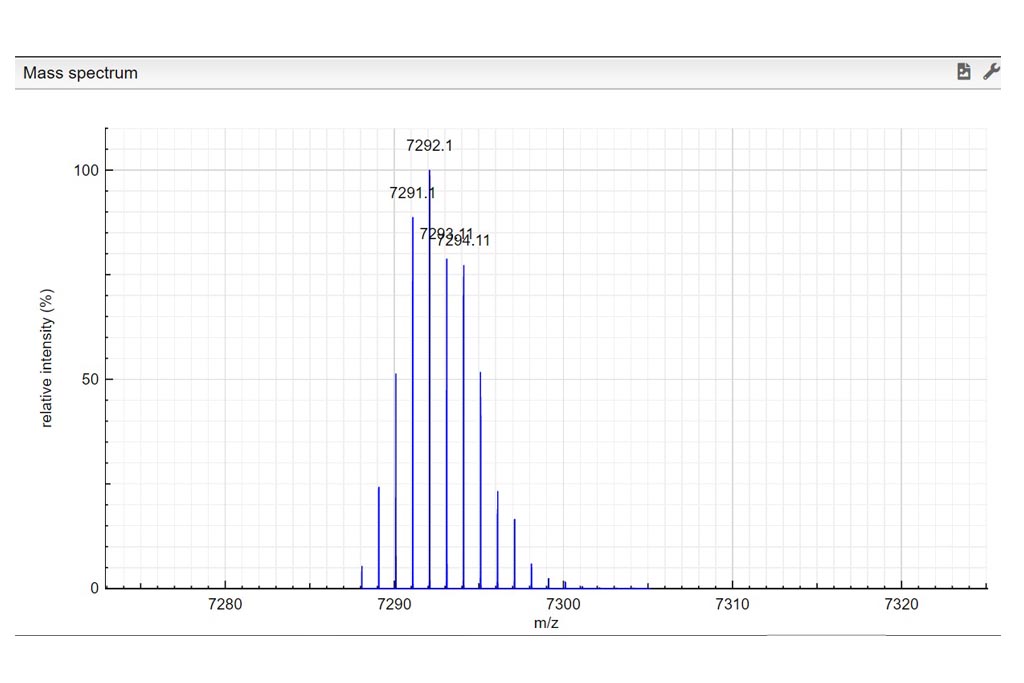
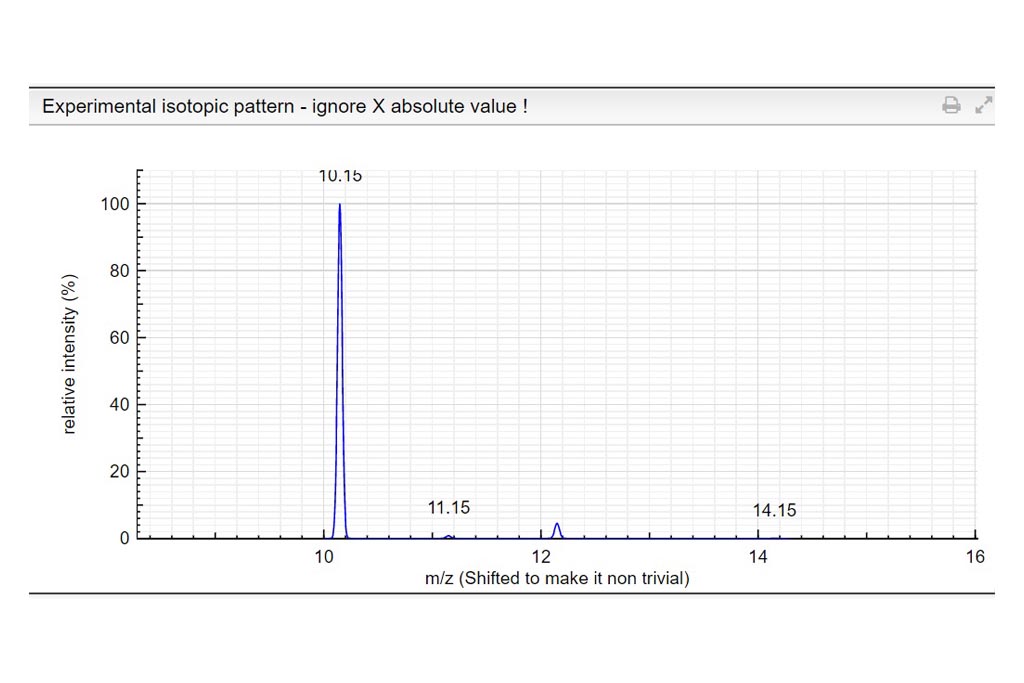
Determine the charge of the observed molecular ion. Depending on the ionization method a molecule can be charged many times. In those examples we only consider positively charged molecules. Based on the theoretical mass spectrum and the difference of mass between isotopologues, determine the charge of the observed molecular ion. Write your answer in the charge column. If it turns green…your answer is correct