STEP 1
Importing Experimental data
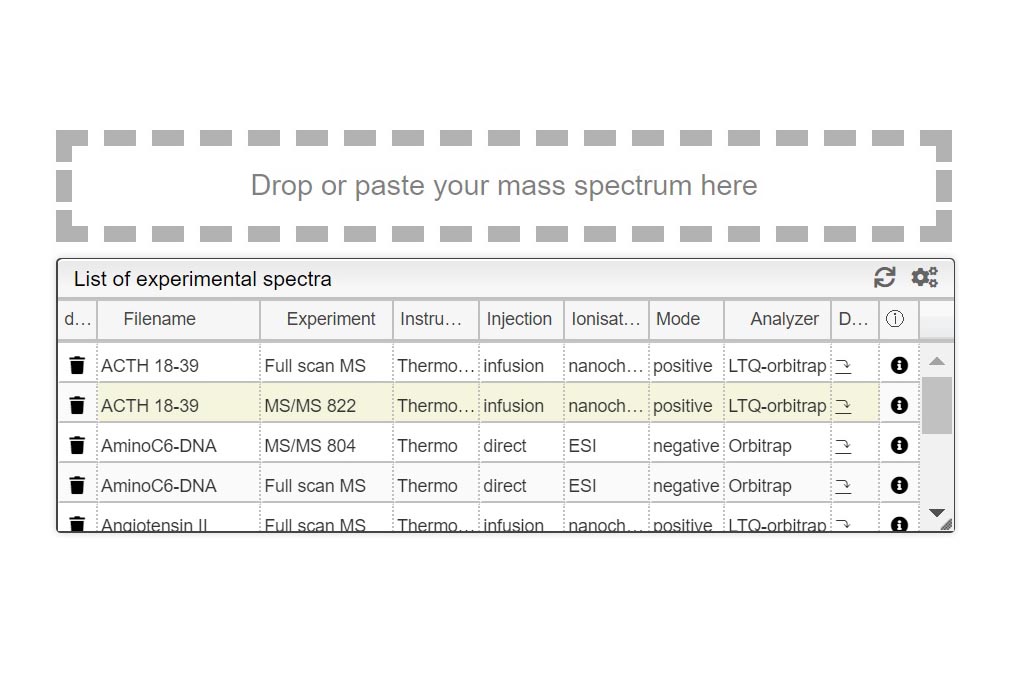
This tool can be either used in a LIMS or stand-alone. In the stand-alone mode you should either drag/drop your experimental spectrum as a tab-delimited text file or copy paste it (CTRL-V) while moving your mouse over the drop zone. The list of available spectra will be displayed in the table and you can simply click on the one you want to display it.